Re-analysis of leukemia data with causalRisk
leukemia.RmdIntroduction
The data come from a multicenter trial of leukemia patients consisting of 137 individuals who were prepared for bone marrow transplants under a radiation-free regimen at four medical centers. Allogenic bone marrow transplants occurred between March 1, 1984 and June 30, 1989. Patients were followed until death or administrative censoring 5 years after transplant.
The goal of the analysis is to estimate the effect on death of a hypothetical time-varying intervention that could prevents graft-versus-host disease (gvhd) from occurring compared to the natural course (no intervention). Baseline covariates, measured at the time of transplant, include age (age), sex (male), leukemia type (acute lymphocytic or acute myeloid leukemia) (acute_lymph_leuk), wait time from leukemia diagnosis to transplantation (waittime), and cytomegalovirus immune status (yes or no) (cmv). During follow-up, assessments were made to determine if and when platelets returned to normal range (platelets), if and when patients experienced leukemia relapse (relapse), and if and when patients developed graft-versus-host disease (gvhd).
Several baseline characteristics of the patients are provide below:
leukemia %>% dplyr::filter(Start == 0) %>%
make_table1(vr=c("Sex","CMV", "age"))| Characteristic1 | |
|---|---|
| Sex | |
| Female | 57 41.61 |
| Male | 80 58.39 |
| CMV | |
| Absent at baseline | 69 50.36 |
| Present at baseline | 68 49.64 |
| age, Mean (SD) | 28.36 (9.56) |
| 1 All values are N (%) unless otherwise specified | |
Unadjusted risk of death
A standard Kaplan-Meier estimator shows the unadjusted 5-year risk of death was close to 60%.
models = specify_models(identify_interval(Start, Stop),
identify_subject(id),
identify_censoring(EndofFollowup),
identify_outcome(death))
km = estimate_ipwrisk(leukemia, models, label=c("Unadjusted risk"),
times = seq(0, 365*5, 5), nboot = 250)
plot(km) + ylab("Cumulative Risk") + xlab("Time in days")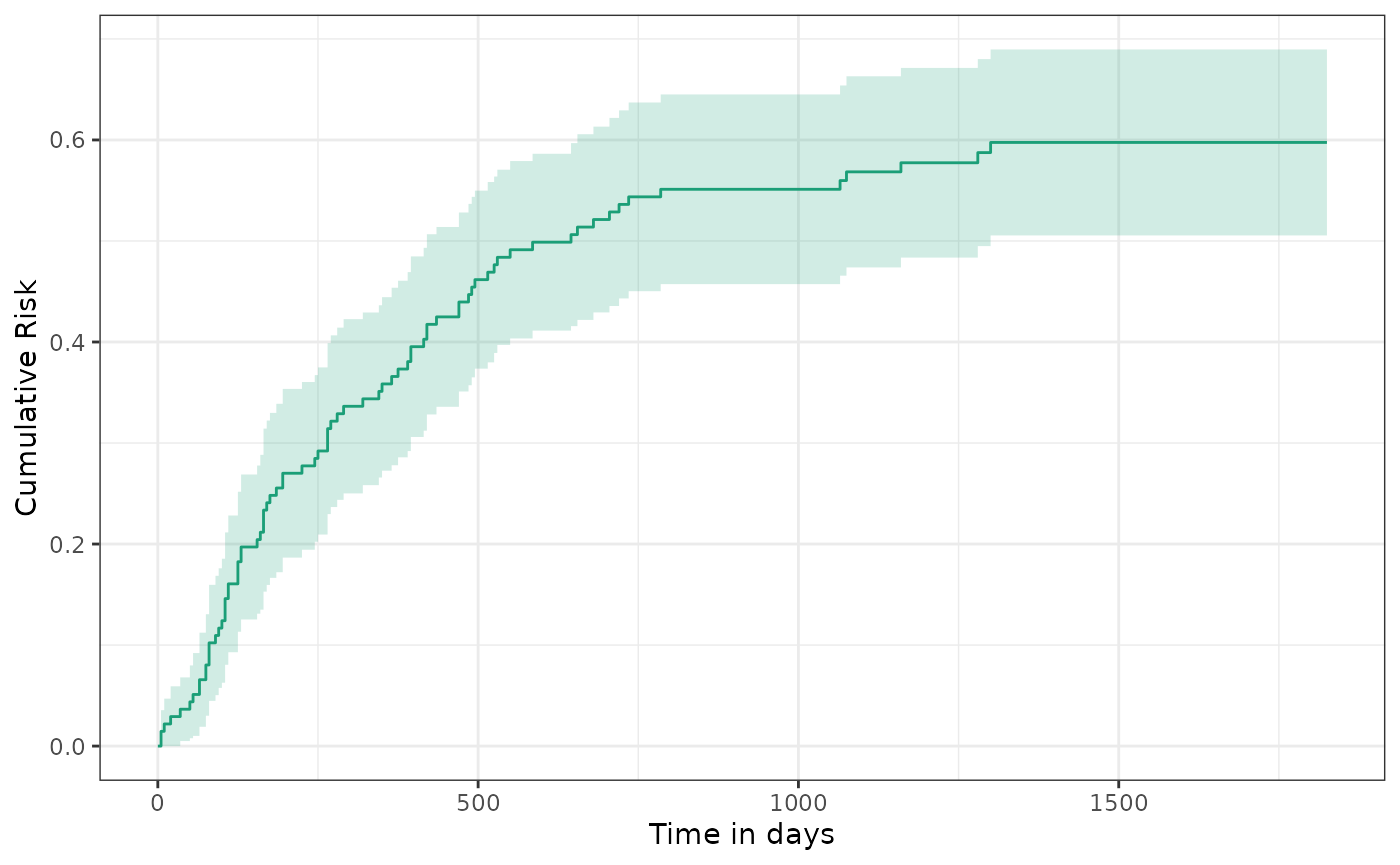
make_table2(km, risk_time = 5*365, pt_scale = 365)The natural course: risk of death adjusted for dependent censoring
To estimate the risk under the natural course, we must control for informative censoring. We consider all baseline and time-varying covariates to be sufficient to assume conditional exchangeability between those censored and those remaining under follow-up. We observed little evidence of informative censoring with the estimated risk remaining close to 60%.
natural_course = km %>%
update_censoring(EndofFollowup,
new_formula = ~age + sex + cmv + acute_lymph_leuk + waitdays + gvhd +
relapse + platelets) %>%
update_label("Risk under the natural course") %>%
re_estimate()
plot(km, natural_course, scales = "fixed") + ylab("Cumulative Risk") + xlab("Time in days")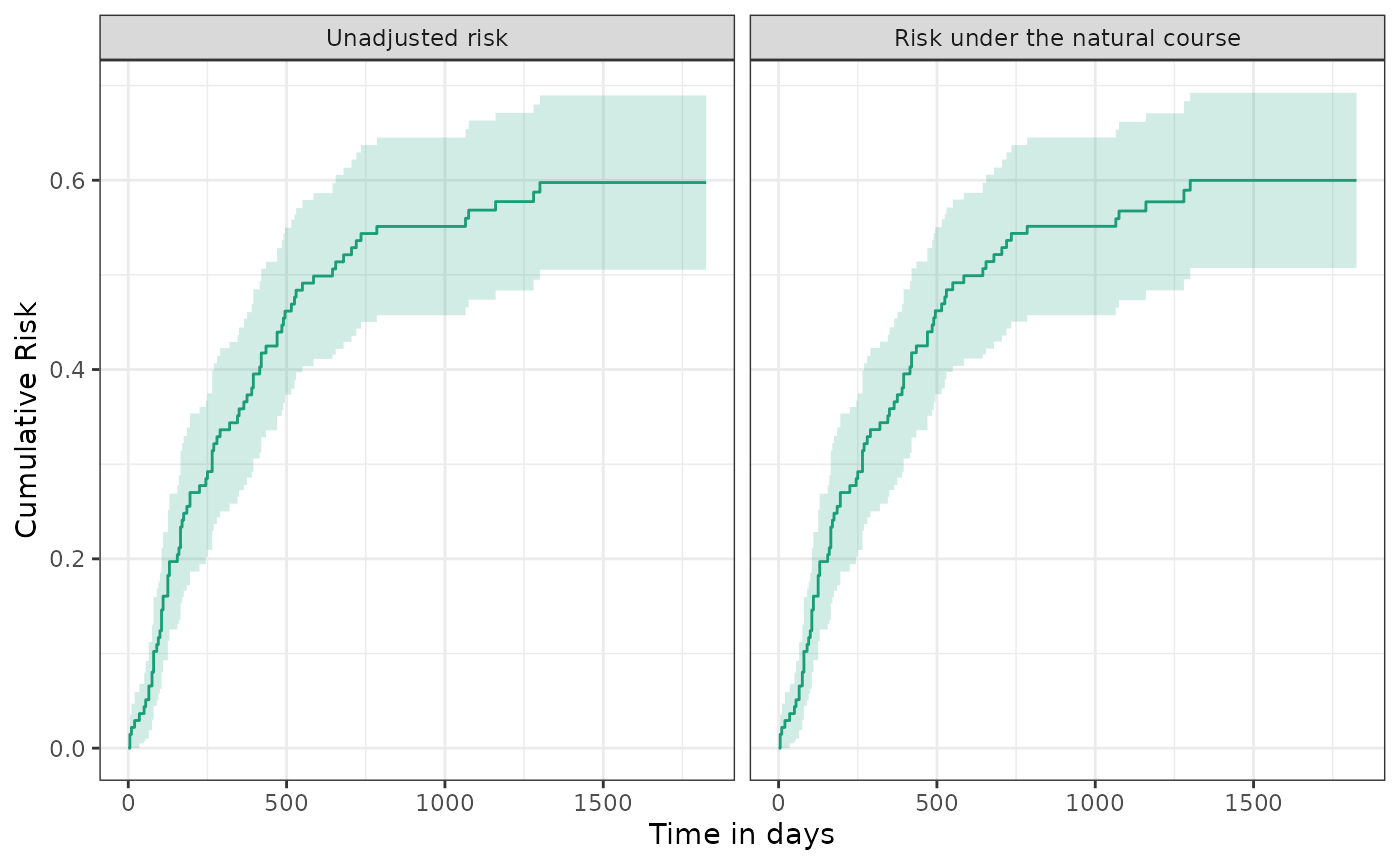
make_table2(km, natural_course, risk_time = 5*365, pt_scale = 365)Counterfactual risk of death if graft-versus-host disease could be prevented
Finally, we sought to estimate the effect of a hypothetical intervention that would prevent all graft-vs-host disease. To do this, we artificially censor patients who develop graft-versus-host disease and adjust for potentially informative censoring through inverse-probability of censoring weights. As is evident, the hypothetical intervention had little effect, with the risk remaining close to 60%.
counterfactual = natural_course %>%
update_censoring(EndofFollowup,
new_formula = ~ . - gvhd) %>%
update_censoring(gvhd_time) %>%
update_label("Risk if GVHD were prevented") %>%
re_estimate()
plot(km, natural_course, counterfactual, ncol = 3, scales = "fixed") +
ylab("Cumulative Risk") + xlab("Time in days")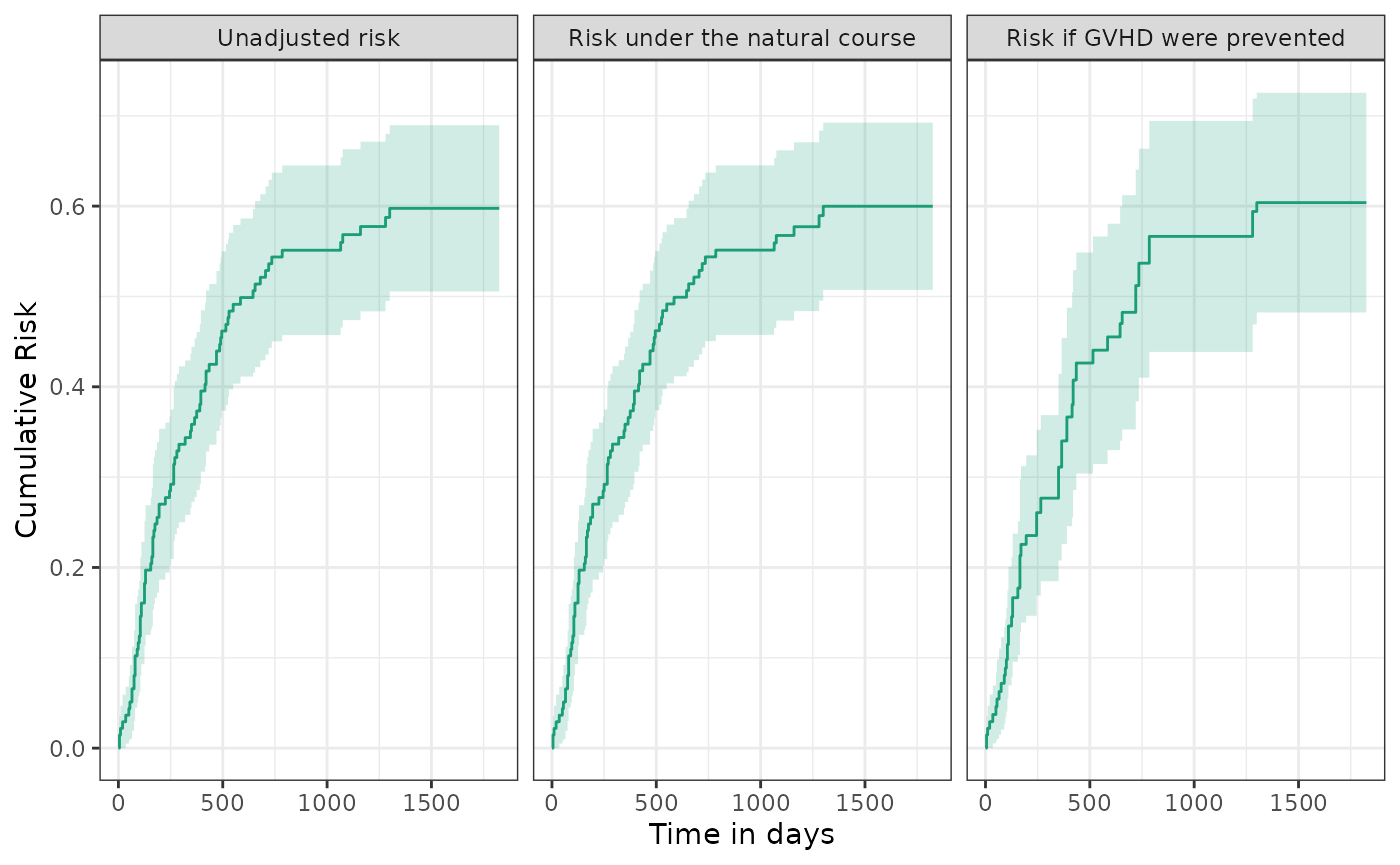
make_table2(km, natural_course, counterfactual, risk_time = 5*365, pt_scale = 365)